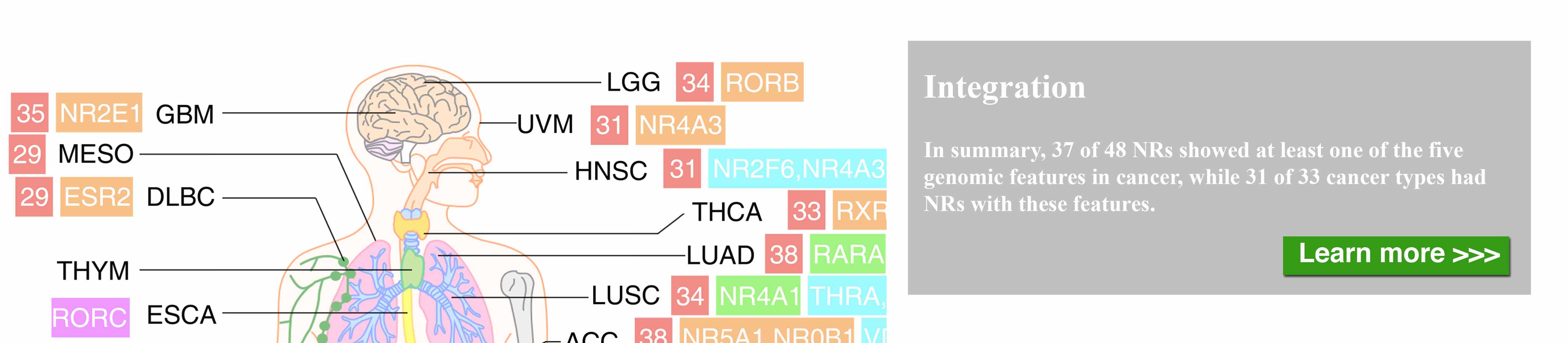
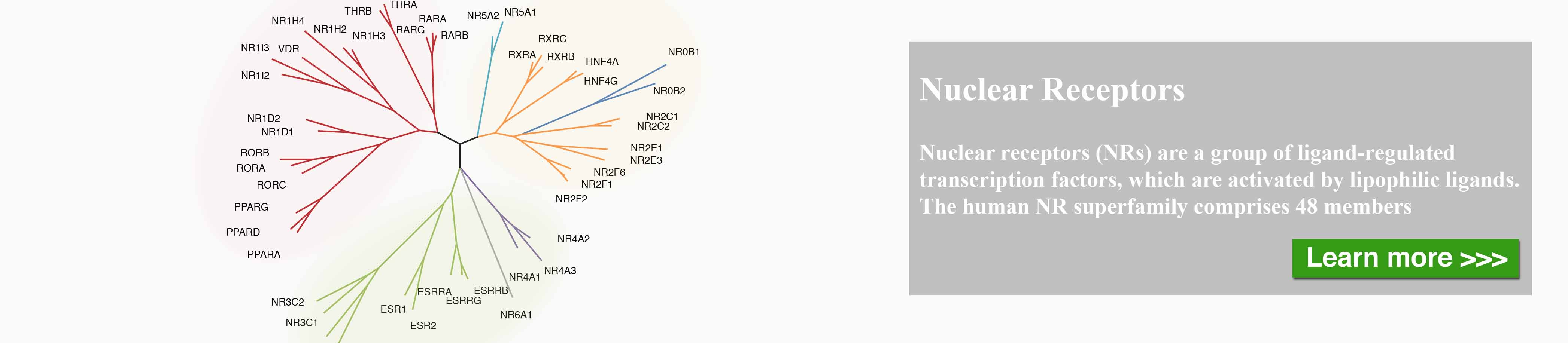
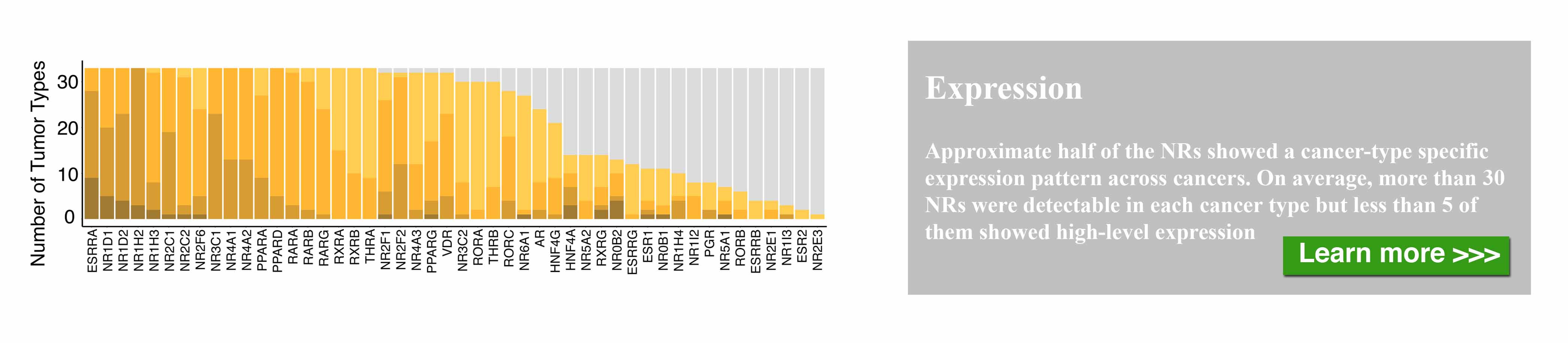
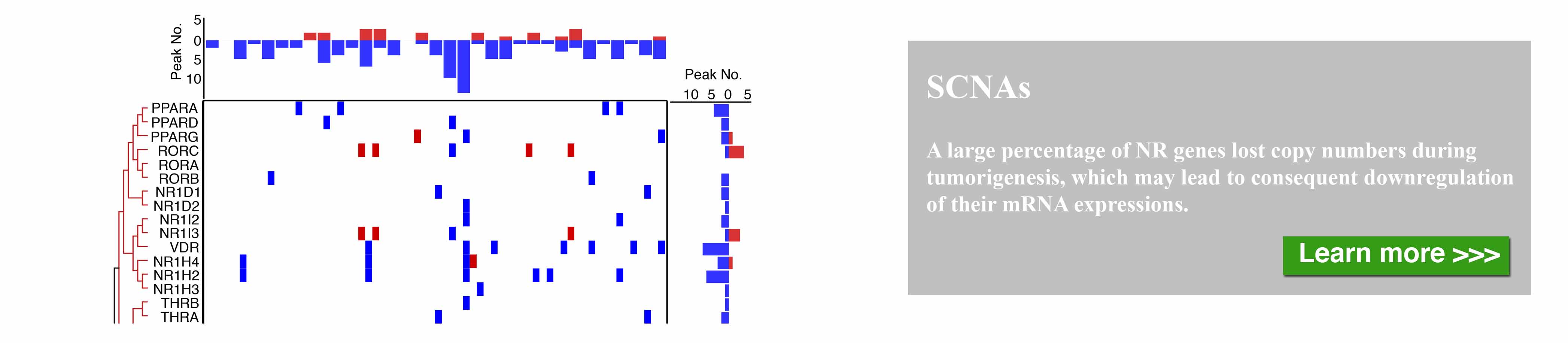
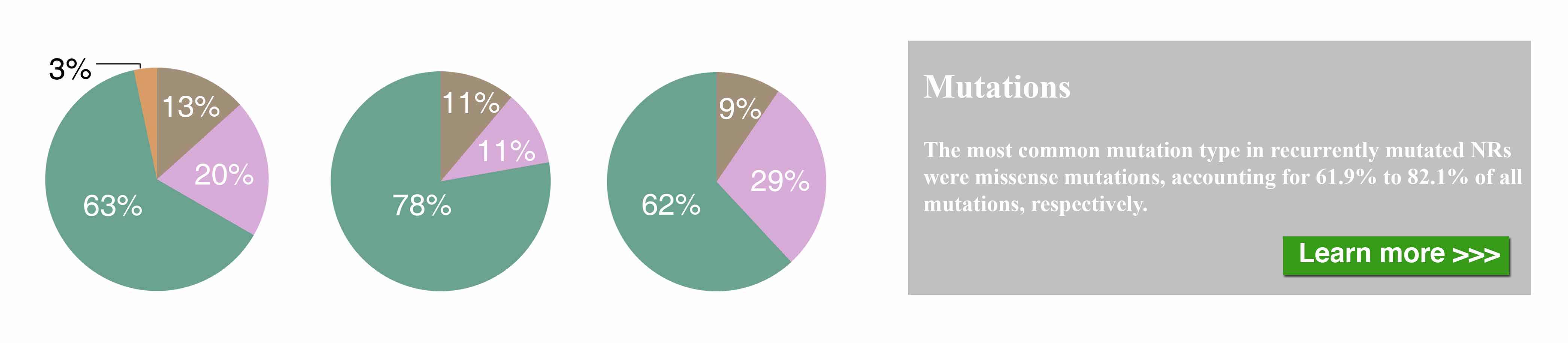
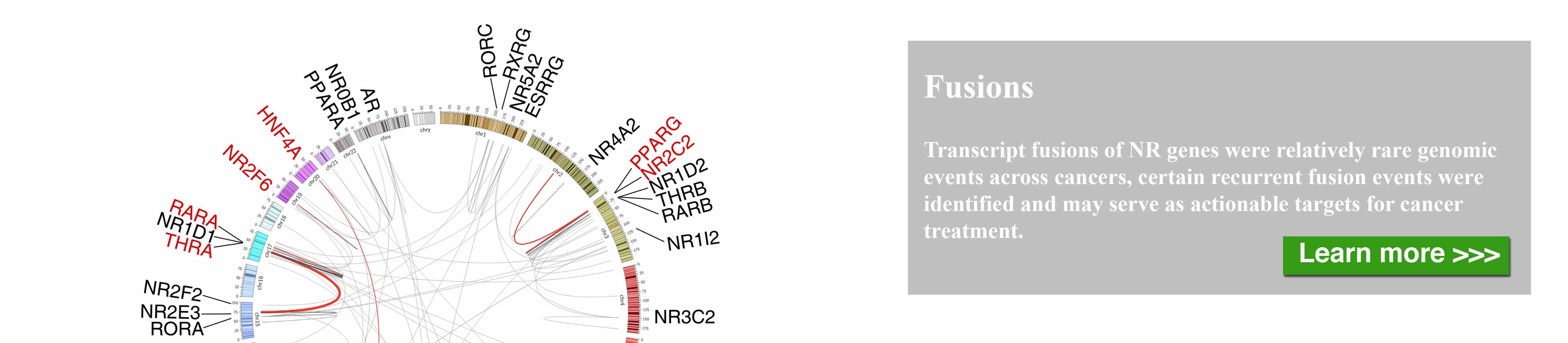
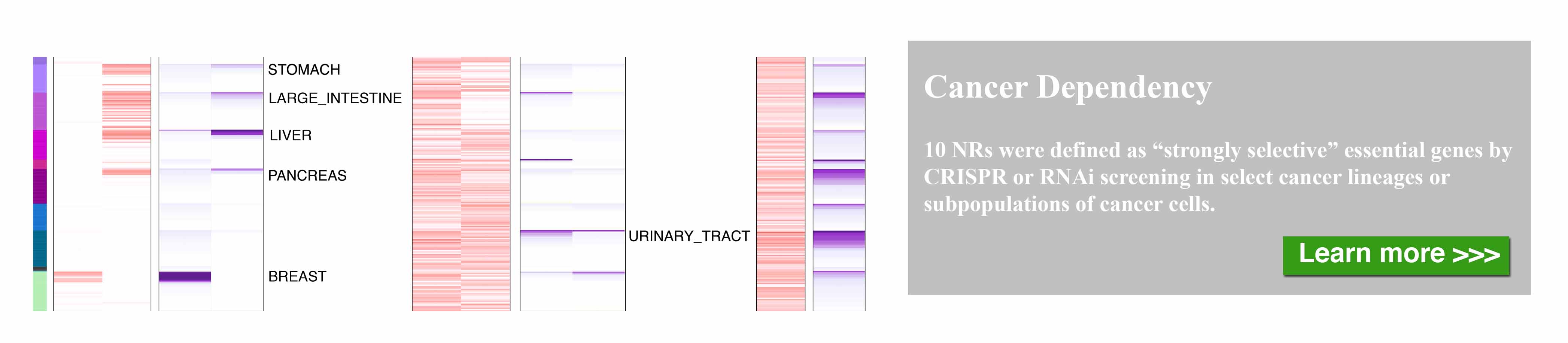

The nuclear receptor (NR) superfamily is one of the major druggable gene families, representing approximately 13.5% of approved drugs. Certain NRs, such as estrogen receptor (ER) and androgen receptor (AR), have been well demonstrated to be functionally involved in cancer and serve as informative biomarkers and therapeutic targets in oncology. However, a comprehensive spectrum of NR dysregulation across cancers remains uncharacterized. Through computational integration of genetic, genomic, and pharmacologic profiles, we characterized the expression, recurrent genomic alteration, and cancer dependency of NRs at a large scale across primary tumor specimens and cancer cell lines. Expression levels of NRs were highly cancer-type specific and globally downregulated in tumors compared to corresponding normal controls. Although the majority of NRs showed copy number losses in cancer, both recurrent focal gains and losses were identified in select NRs. Recurrent mutations and transcript fusions of NRs were observed in a small portion of cancers, serving as actionable genomic alterations. By a combination of large-scale CRISPR and RNAi screening profiles, 10 NRs were defined as strongly selective essential genes for cancer cell growth. Their growth dependencies were significantly correlated with their expression or genomic alterations in a subpopulation of tumor cells. Our comprehensive characterization of NRs across cancers may facilitate the identification and prioritization of potential biomarkers and therapeutic targets, as well as the selection of patients for precision cancer treatment.